Further investigations of the diversity of the Glyceriformia (Polychaeta: Goniadidae and Glyceridae) from West African shelf areas
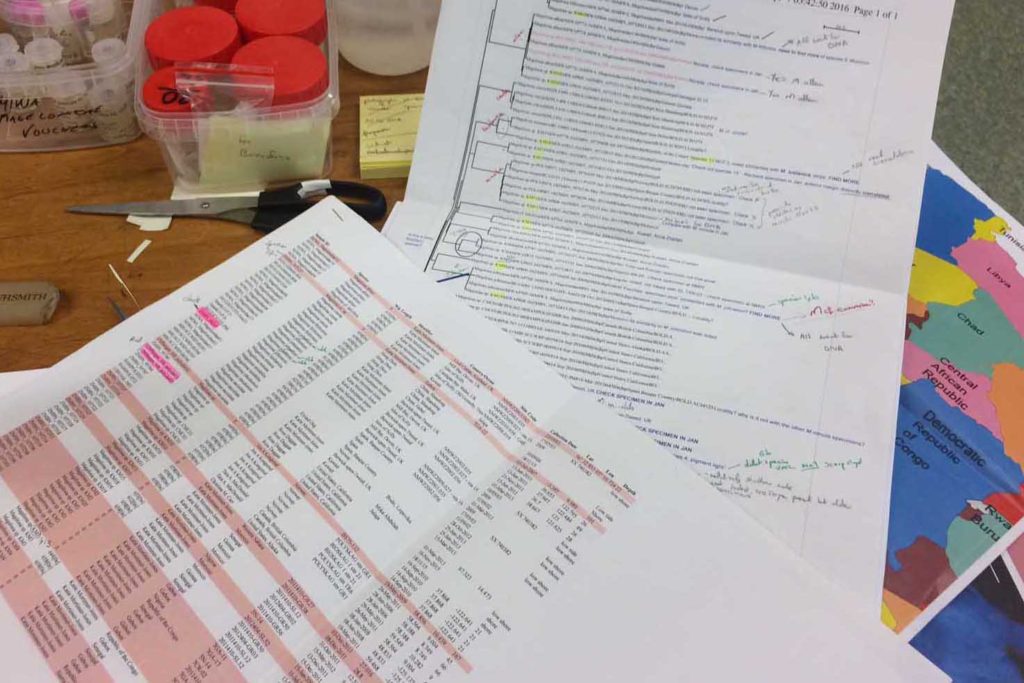
For the past three weeks, Lloyd, who is a Senior Environmental Scientist at Envaserv Research Consult in Ghana, has been visiting us to continue work on the project we have running on the Glyceriformia in the MIWA-material. This project was initiated when Lloyd and Willams were here visiting in November 2015. They worked closely together with Tom, using the available literature to identify and select animals for barcoding, and to get a feel for the diversity of the group.
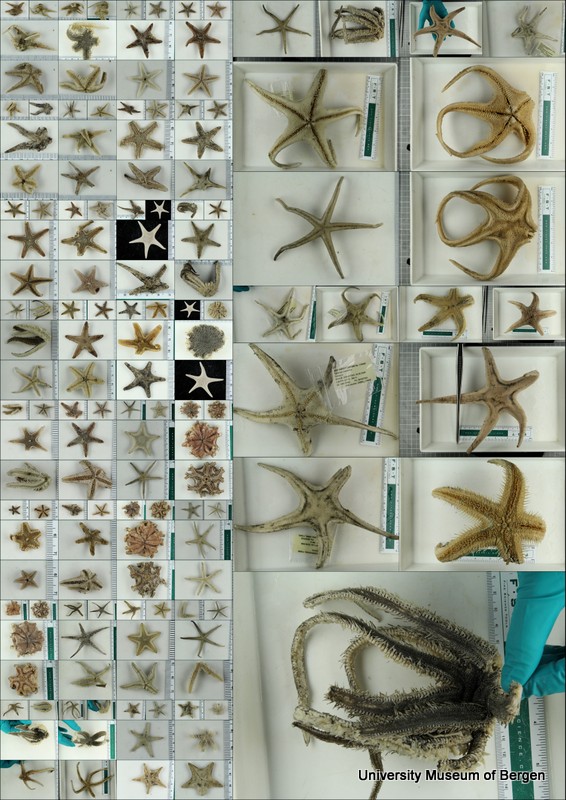
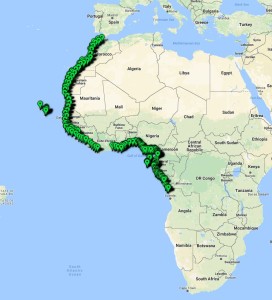
Similar projects on other polychaete families have taught us that the current knowledge of species diversity and distribution of the region is not comprehensive – there are more species than what is currently described in the literature, so we attempted to barcode representatives of all the morphologically distinct groups – it’s highly likely that material contains species that are new to science.
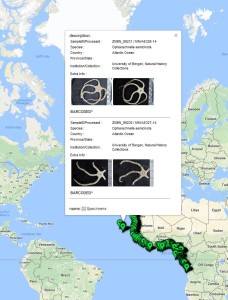
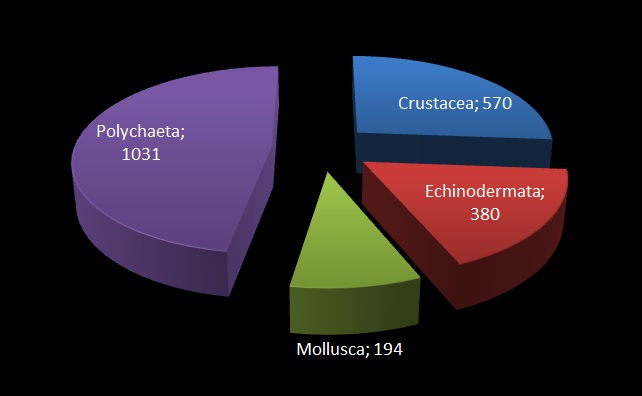
From this work, 19 “species” were identified based on morphology, and several representatives for each were selected for genetic barcoding: A voucher specimen is selected, photographed, and tissue sampled. The tissue sample is sent to the CCDB lab in Canada for sequencing, whilst the photo and metadata (such as where the animal was collected, who has identified it, at which institution is the voucher specimen stored etc.) is uploaded to the international database named BOLD – Barcode of Life Datasystems, as part of the iBOL (international barcode of life) initiative.
Unfortunately, only 50% of the specimens we tried to  barcode resulted in a barcode sequence, but even so the DNA indicated 21 genetically distinct groupings (“BINs”).
barcode resulted in a barcode sequence, but even so the DNA indicated 21 genetically distinct groupings (“BINs”).
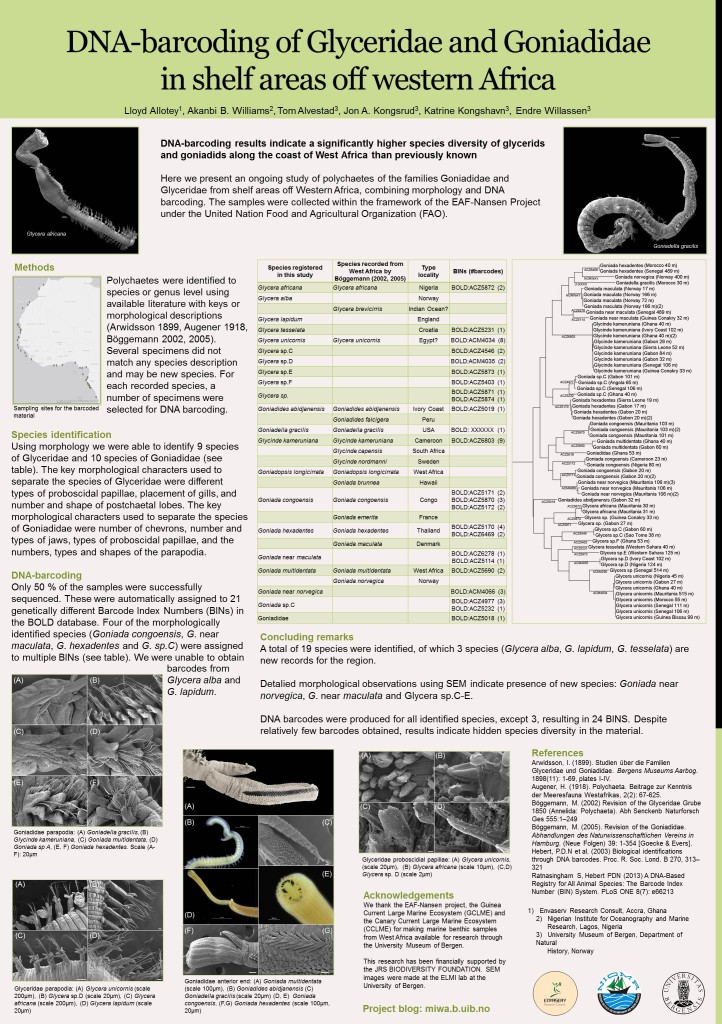
Last summer we made a poster summarizing the work we had done, and presented it as a poster at the 12th International Polychaete Conference in Wales in August 2016.
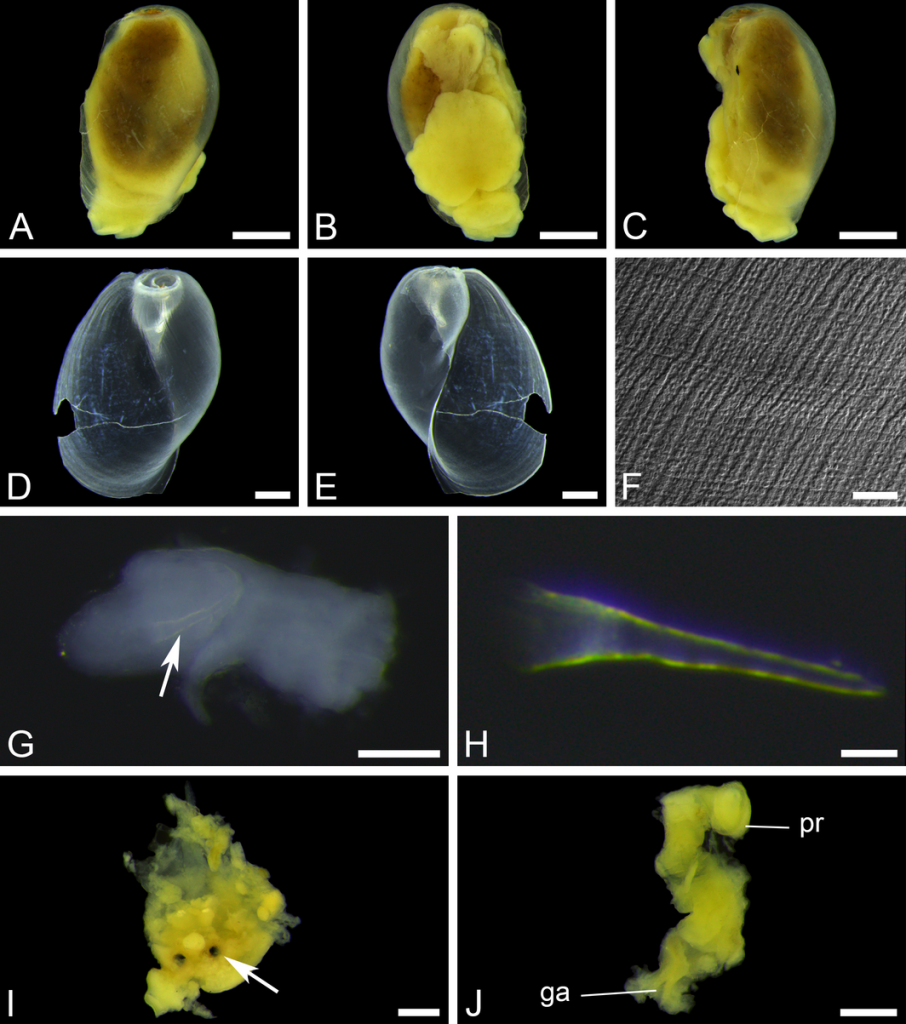
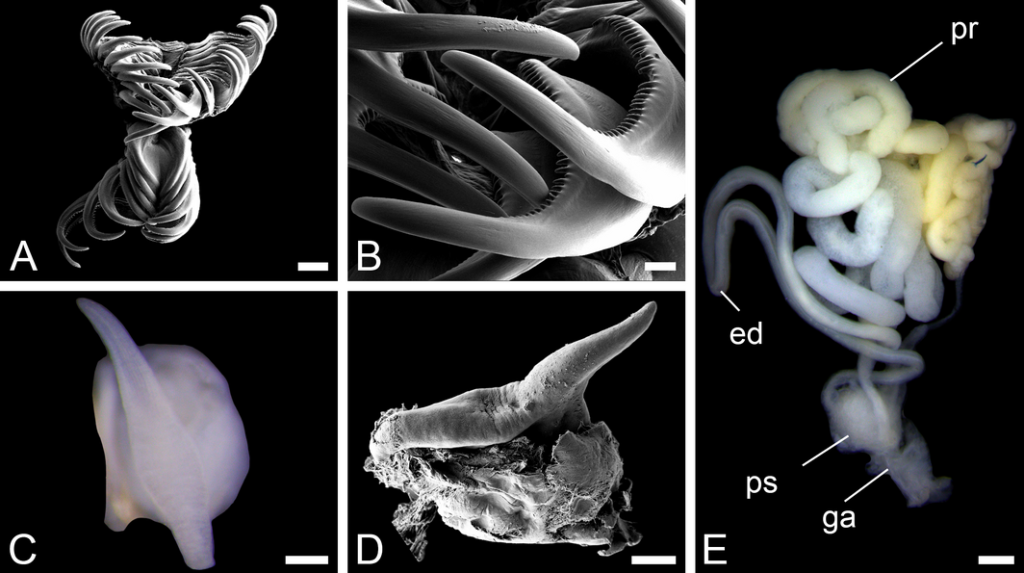
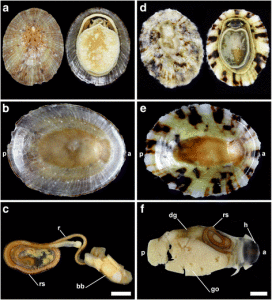
We’ve continued the work, both with sorting more samples, thus making more material available for studies, and by doing detailed examinations of the specimens we have. This is done using both regular light microscopy and scanning electron microscopy (SEM).
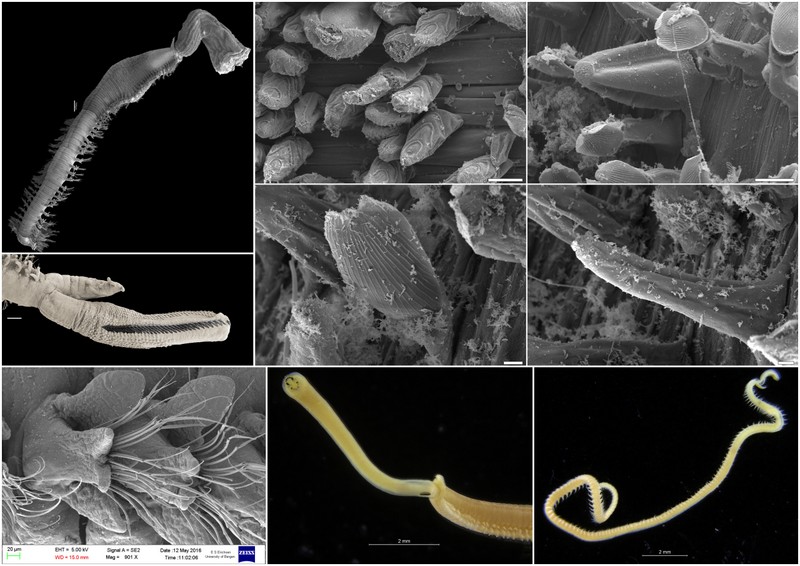
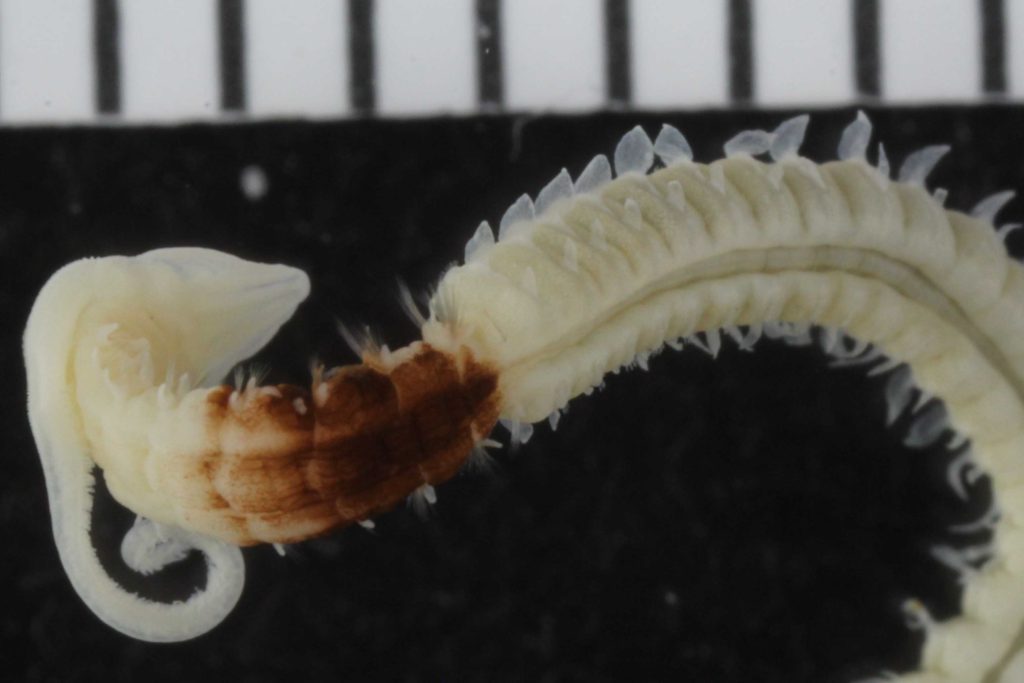
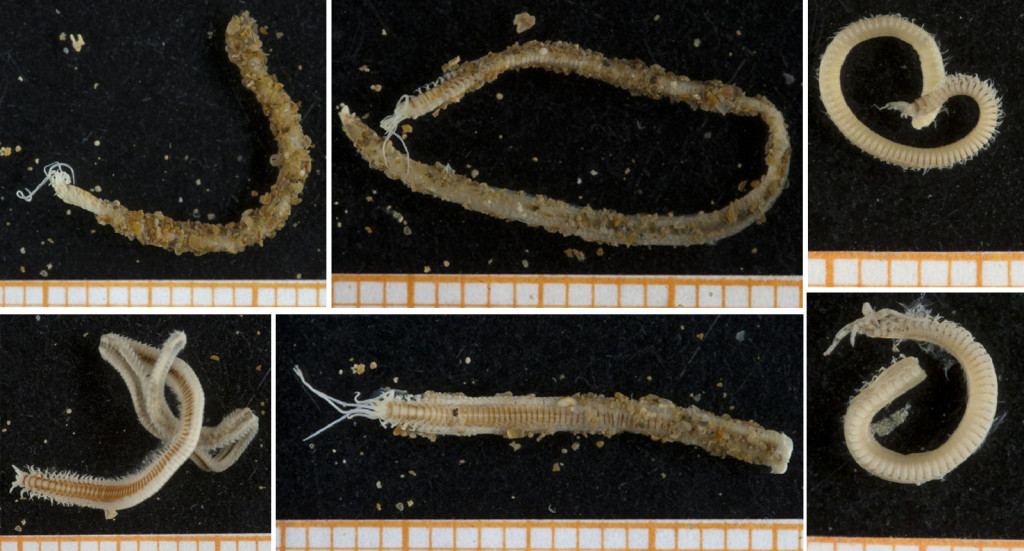
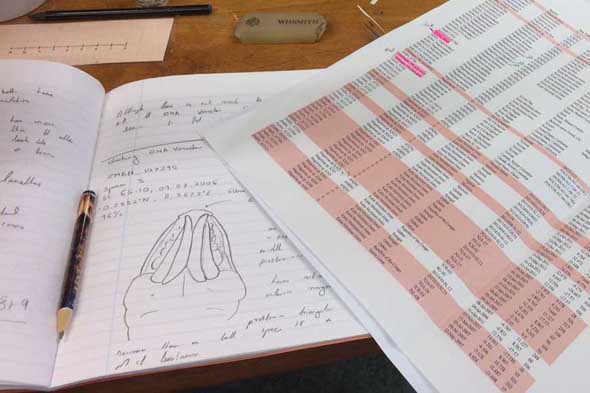
Some of the characters that needs examination – overall morphology, jaw structures, parapodia and bristles, and the papillae on the proboscis
During his most recent stay, Lloyd continued the work with identifying animals and selecting specimens that we will submit for barcoding – we’ll try to get the next plate sent in by the end of the month, and hope for a high(er) success rate and further insights in the diversity of the Glyceriformia of the region.
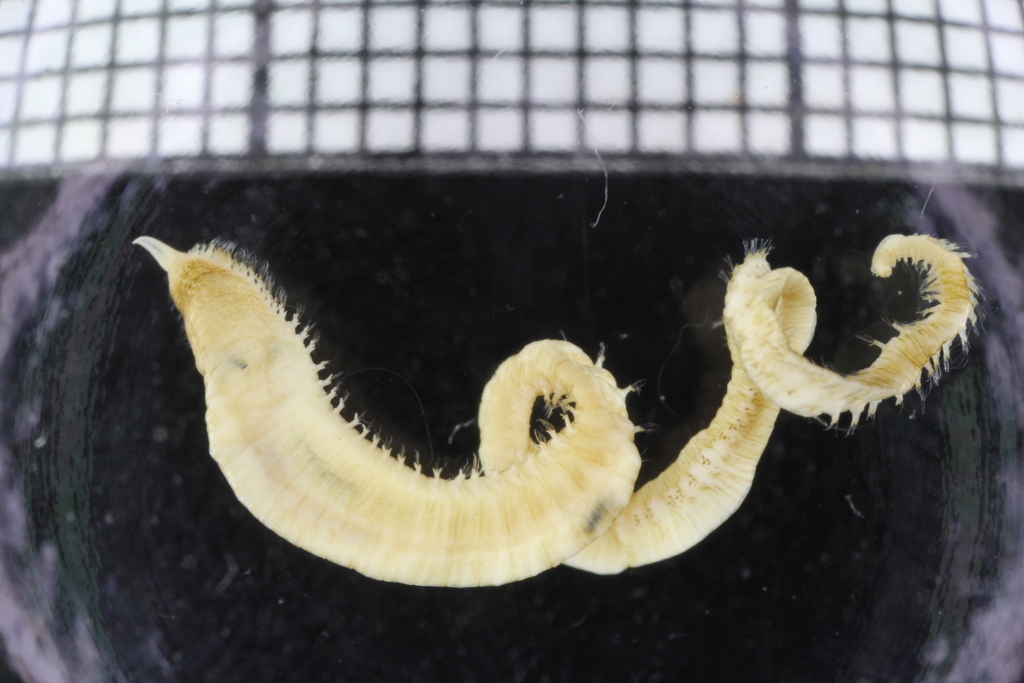
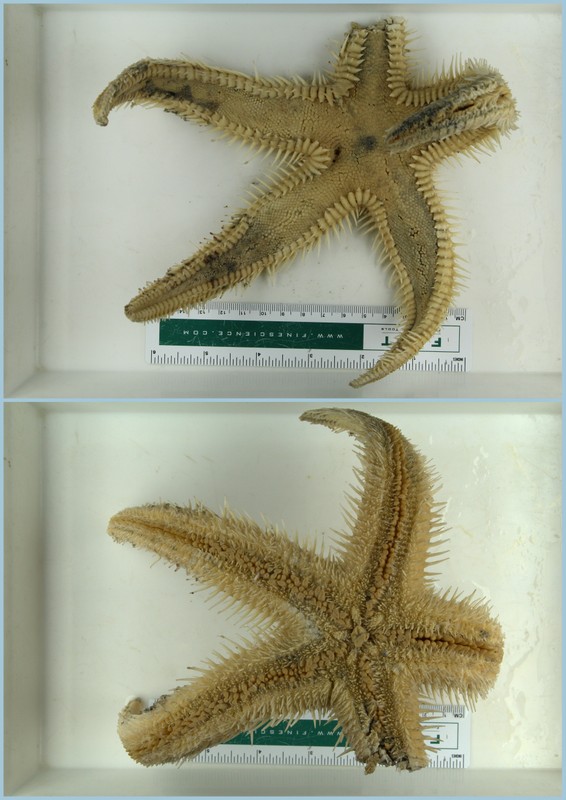
- Glycera unicornis from Guinea Bissau
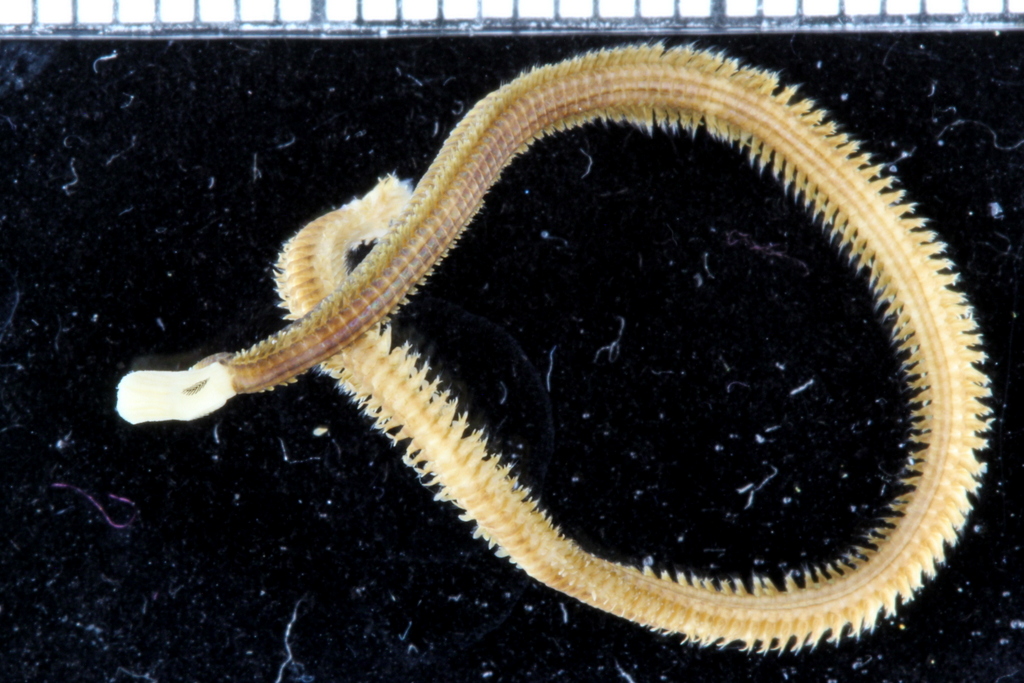
- Glycera sp from Morocco
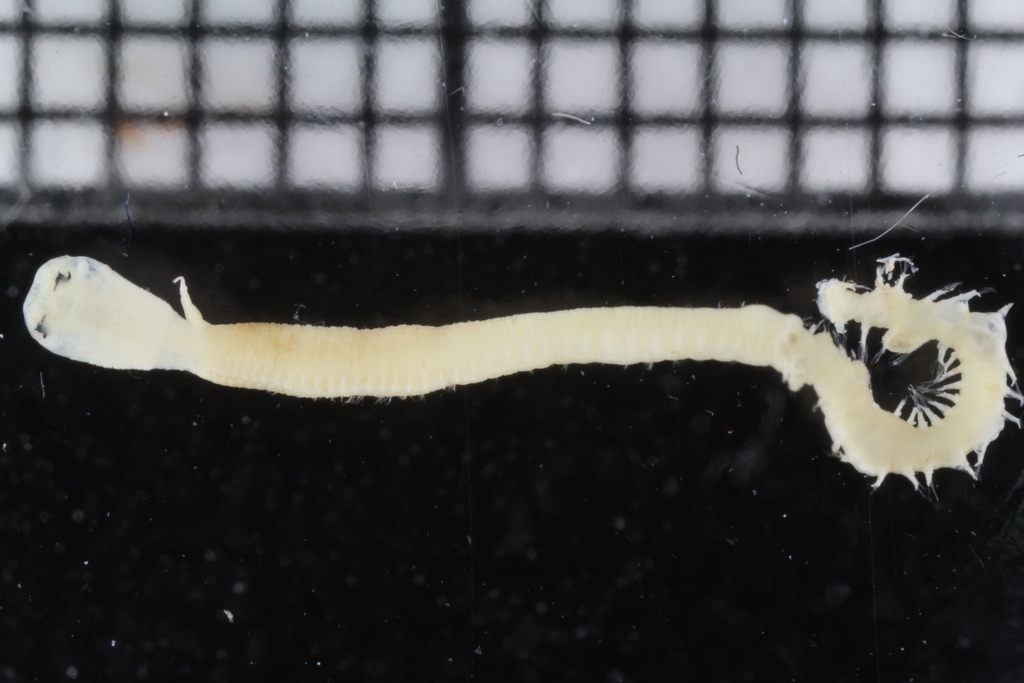
- Goniada congoensis from Senegal
- Glycera sp C from West Africa
Thank your for visiting, and for all your hard work, Lloyd! We hope to see you again soon.