One of the topics covered in depth during the workshop is the selection of specimens and the preparation of tissue-samples. These will to be sent for DNA sequencing, and the genetic sequence will then be included in the Barcode of Life Data Systems (BOLD). The aim is to obtain standardized genetic sequences (“barcodes”) for the various taxa that we are working on. The barcode consists of a segment of approximately 650 base pairs of the mitochondrial gene cytochrome oxidase c subunit 1 (COI). You can read more about DNA barcoding on WIKIPEDIA.
There is a very real challenge connected with estimating biodiversity when many of the species are still undescribed, as is the case with many invertebrate species, especially the more obscure and diminutive groups. In such cases, barcoding can serve as a tool in screening for biodiversity, and aid the taxonomists in identifying areas where the taxonomic resolution is low.
We have not yet received any barcodes for our MIWA project, but the project page on BOLD is getting populated with images and geographical information.
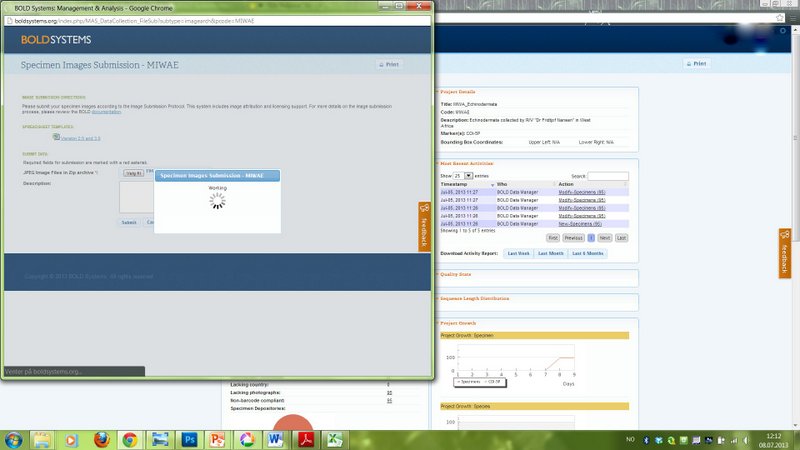
Uploading pictures
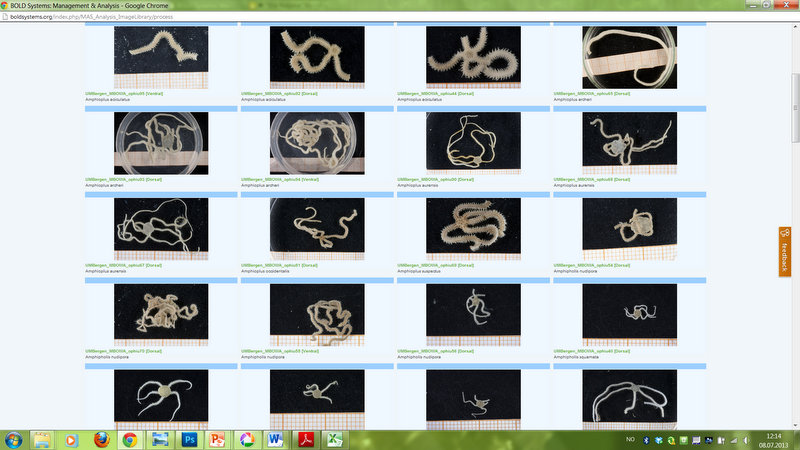
Brittle star image gallery
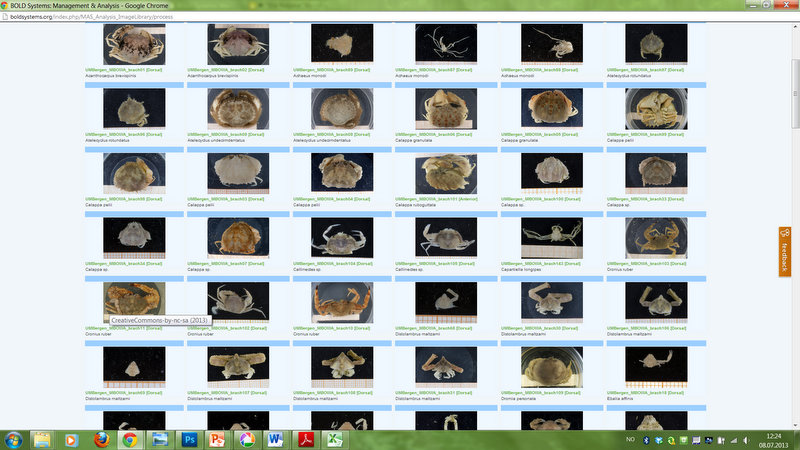
Crab image gallery
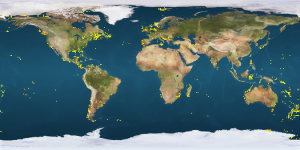
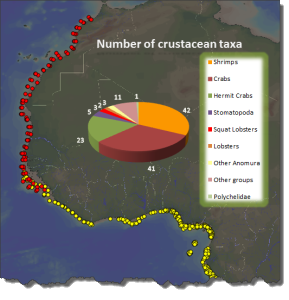
This picture shows a plot of crab species that have been DNA barcoded around the World. Notice the lack of records from West African waters.
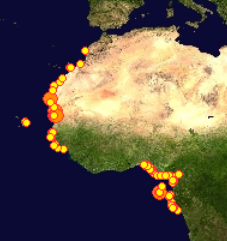
This picture shows how we are presently about to add records of about 60 crab species to the BOLD database.
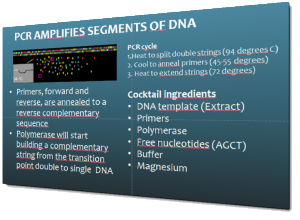

How is it that the polymerase chain reaction (PCR) can make numbers of copies of a piece of DNA? (Slide from an intro to the technology of PCR and Sanger sequencing given by EW.)

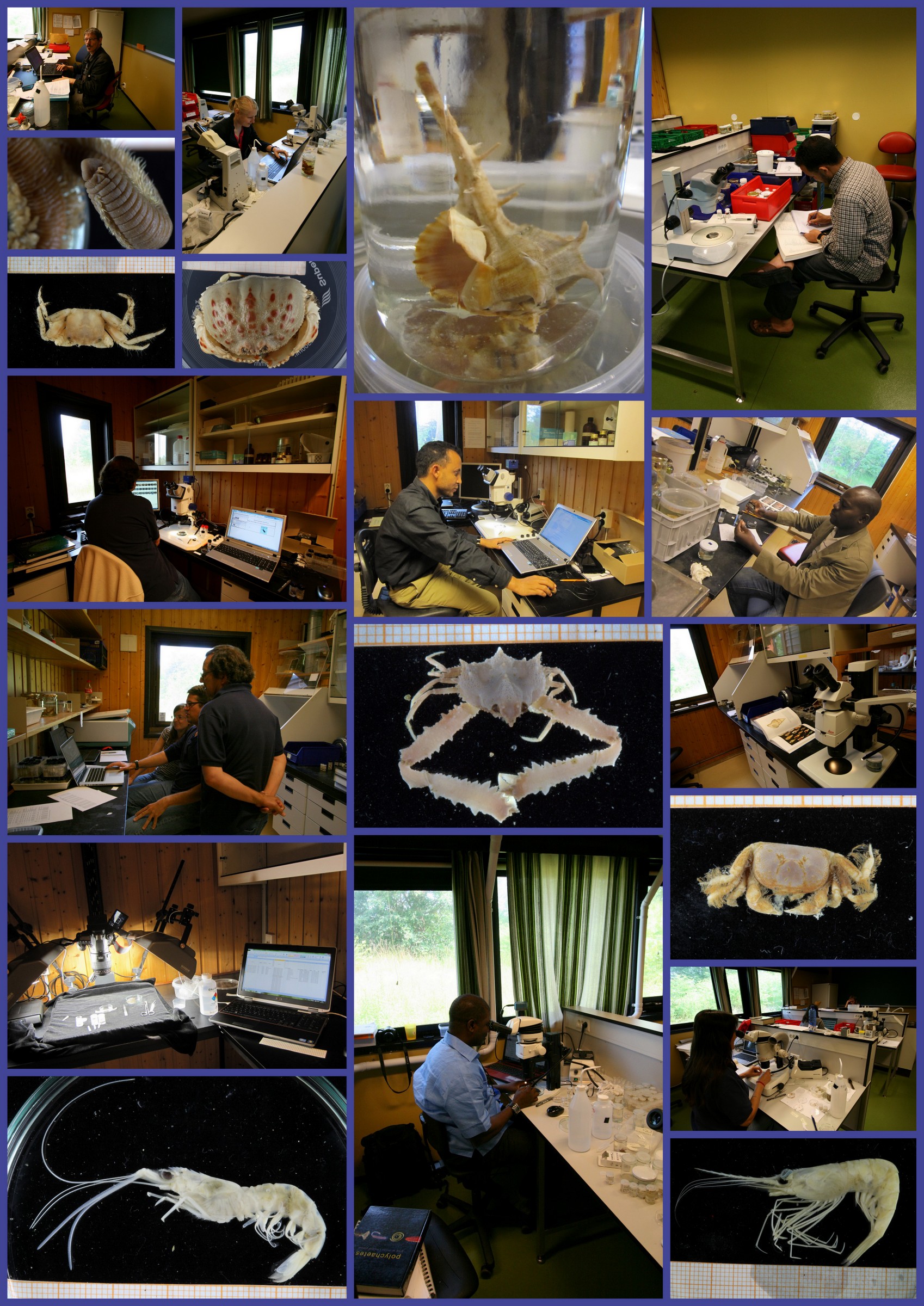
Lecture about the procedure for tissue sampling and recording of data for the BOLD system
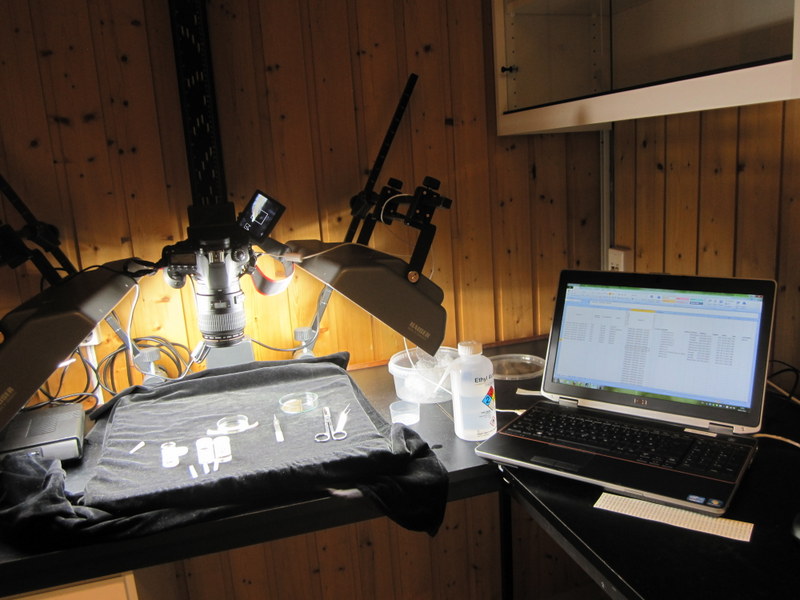

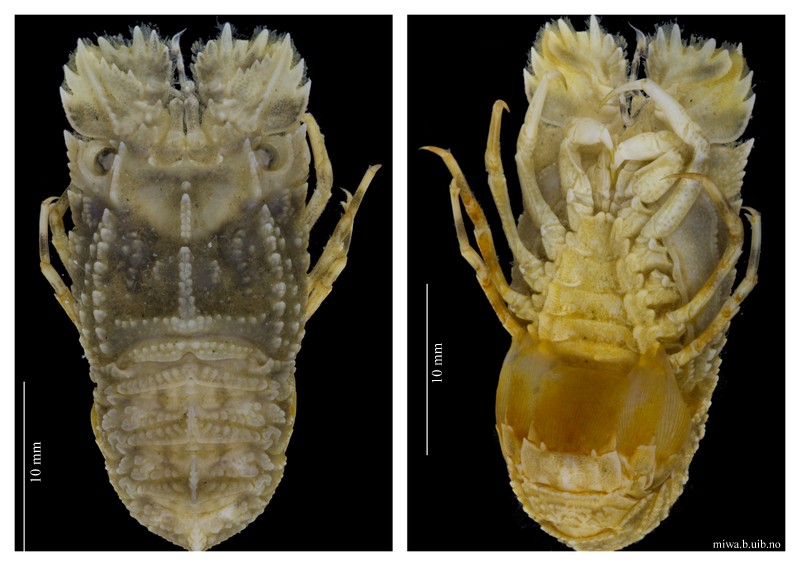
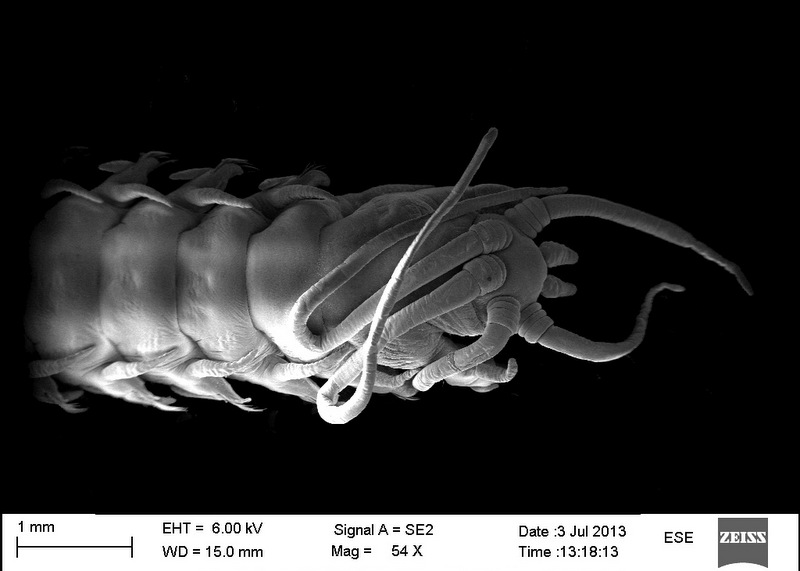
The specimens from which we take the DNA tissue sample is documented through photographs
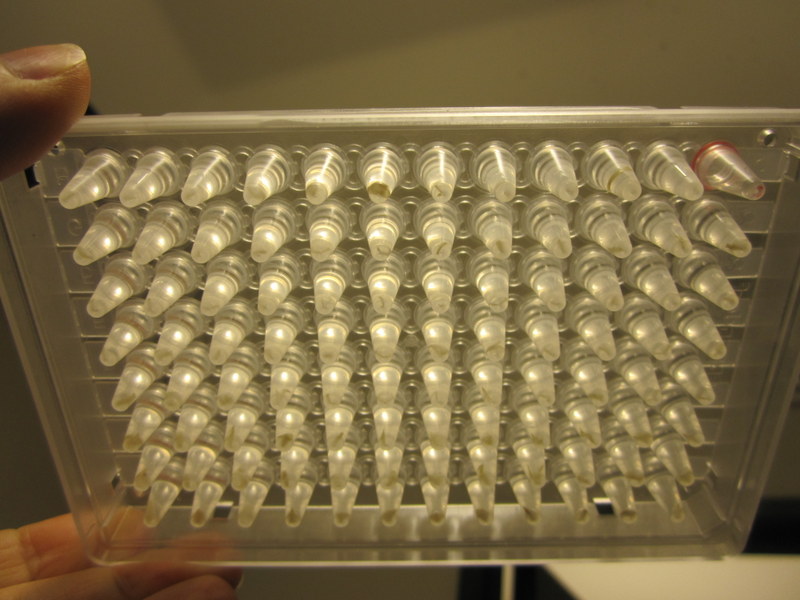

A completed microplate with 95 tissue samples